* Day 1: Preparation of sample
1) Medium preparation for Gluconacetobacter (500ml)
- Glucose - 10g
- Peptone - 2.5g
- Yeast extract - 2.5g
- Na2HPO4 - 1.35g
- Citric acid - 0.75g
- Distilled or tap water - 500ml
- Agar - 7.5g
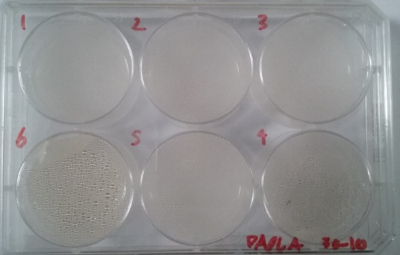
2) Bacteria insolation, a mixture of original bacteria kombucha diluted in distilled water in different proportions.
- 1) 5 parts of original bacteria into 50 parts of distilled water 1/10
- 2) 2 parts of sample #1 into 50 parts of distilled water.
- 3) 2 parts of sample #2 into 50 parts of distilled water.
- 4) 2 parts of sample #3 into 50 parts of distilled water.
- 5) 2 parts of sample #4 into 50 parts of distilled water.
- 6) Distilled water. control sample
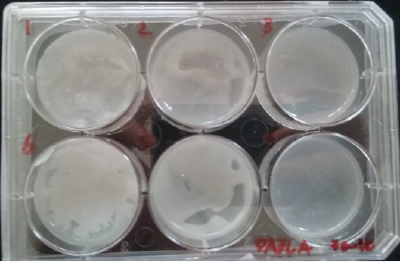
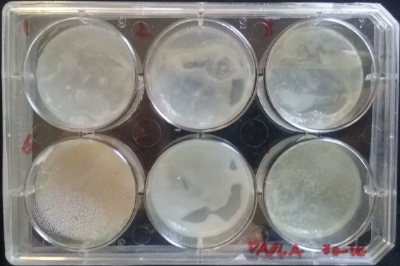
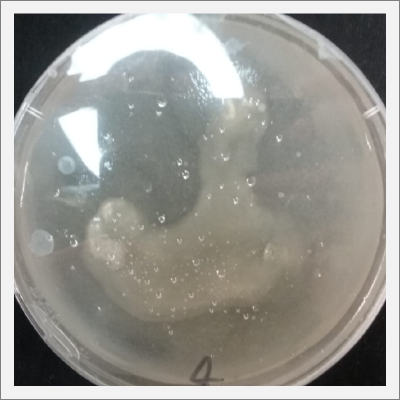
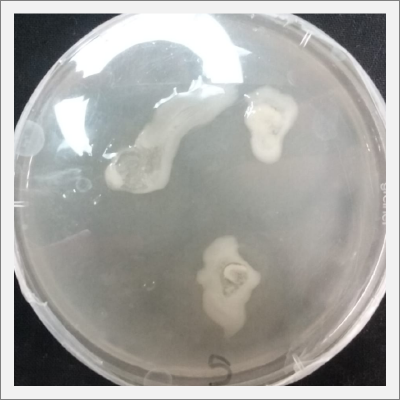
* Day 4: after four days the bacterias seems to be growing, all of them in white color.
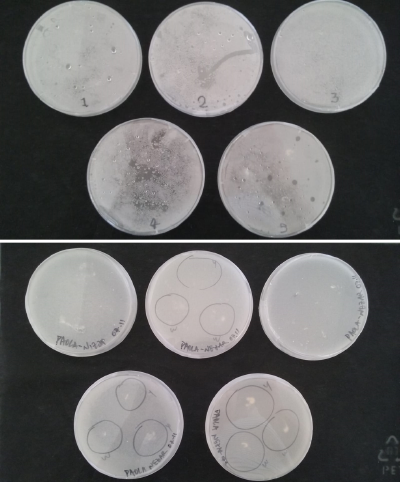
* Day 7: after one week the microorganism (bacteria) that is wanted to be insolated, start to grow, is the white circle spots that can be clearly seen in Petri number 3, 4 and 5.
* Day 8: In order to extract one specimen, 5 new Petri box were made like following;
1) Medium preparation for Gluconacetobacter (100ml)
- Glucose - 2g
- Peptone - 0.5g
- Yeast extract - 0.5g
- Na2HPO4 - 0.27g
- Citric acid - 0.15g
- Distilled or tap water - 100ml
- Agar - 1.5g
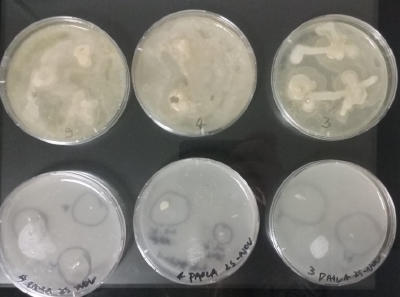
2) Bacteria insolation, from each sample that was made on the first day from 1 to 5 (6 was control water sample) growing bacterias were insolated like following from the original Petri-box samples;
File:Bio-03e.png original Petri-box sample day7
- 1) 3 samples were taken from white spots.
- 2) 1 sample is taken from a yellow spot and two samples are taken from white spots.
- 3) 3 samples were taken from white spots with white dots (the 1 & 2 sample are not dots, they are spots).
- 4) 1 sample is taken from a white dot and 2 samples are taken from yellow/white spots.
- 5) 2 samples are taken from white dots and 1 sample is taken from a yellow spot.
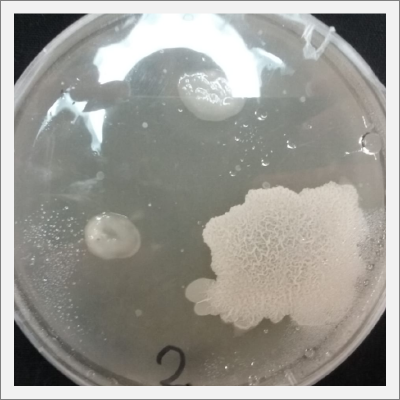
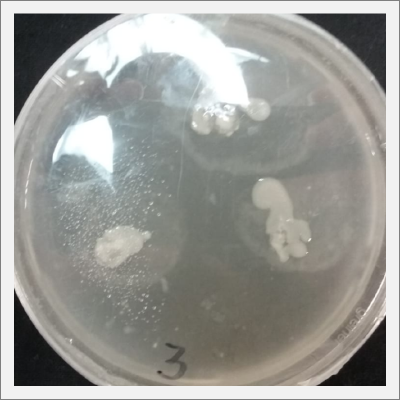
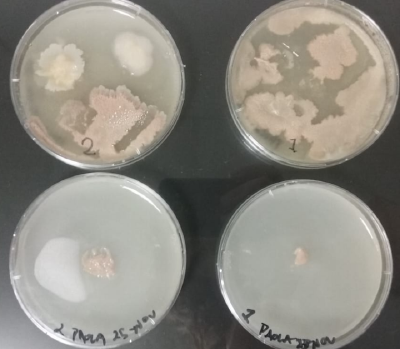
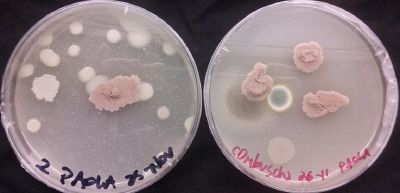
* Day 10: Two days after the transplantation of the samples;
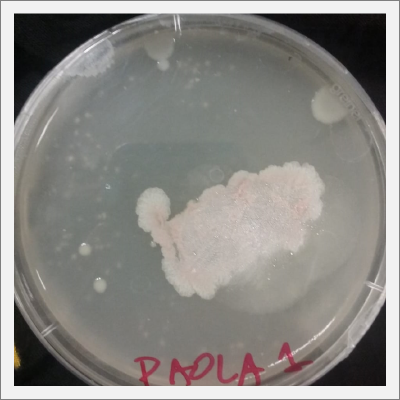
- 1) The three samples have grown into what can be seen in the following pictures.
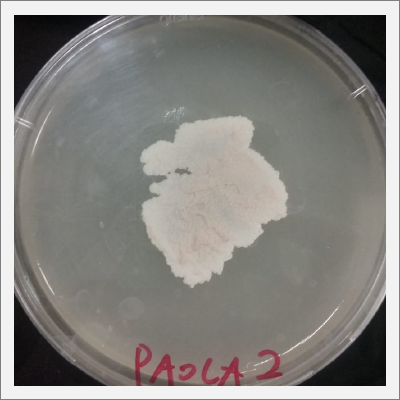
- 2) The sample taken from one of the spots have grown similar to the sample in Petri box #1.
- 3, 4 and 5) The samples that were taken in the other three Petri sample haven shown visible results like the 2 first ones.
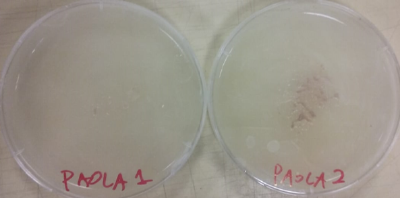
Day 15: I retransplant samples from the 5 Petri dishes in order to try to insolate some new samples that have the same structure as the two first samples, and also transplant some new specimens of the two first samples in new Petri dishes.
Day 18: I take some new clean samples of the two first samples that have an isolated specimen, the Petri dishes 3,4 and 5 didn't grow the desired specimen.
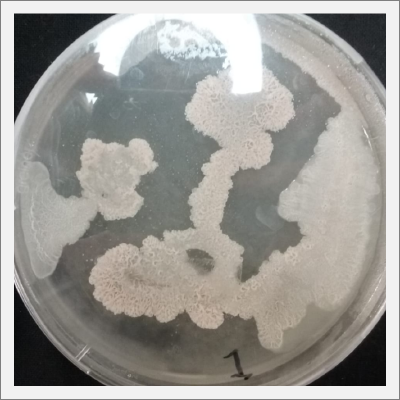
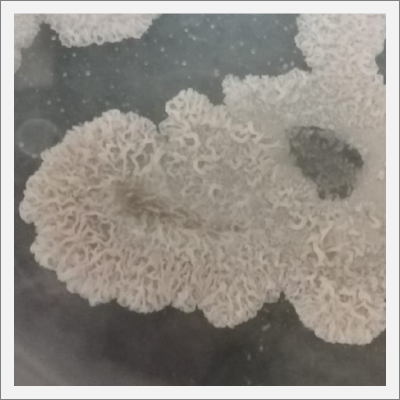
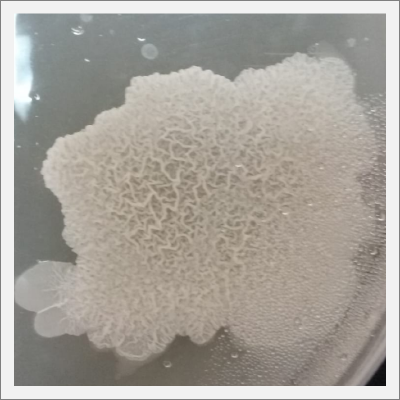
Day 20: The samples are already growing and acquiring its characteristic shape and its rosa and white color.
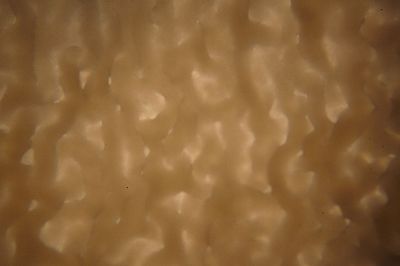
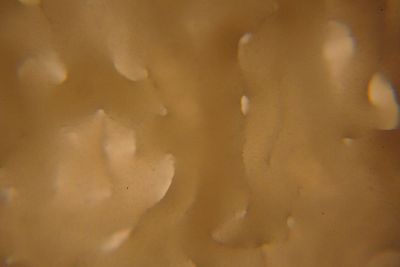
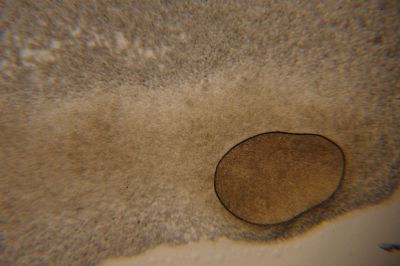
Day 27: I prepare some samples of the specimen in the lens in order to see them under the microscope. The results are shown in the following pictures, with the 40x the structure of the specimen can be seen, with the 100x this structure disappears to the eye, and little brown peaces can be seen.
New specimens are made in order to have some new clean samples.
Day 30: one of the samples taken have inside it penicillin, none of the samples before hat it before. This is a subject of study, in order to see how the isolated specimen behaves in presence of this other specimen, also there's another specimen that had contaminated all of the petri disch of one of this samples before. For the moment it can be seen, that both specimens live one next to the other without a problem.
POSIBLE SPECIMEN
Clonostachys sp.
Fusarium sp.
Acremonium masseei
Acremonium murorum
Talaromyces minioluteus
https://pdfs.semanticscholar.org/4d16/1cea571a604af97d9b05f23a3127eab92a83.pdf
https://www.interdisciplinary-laboratory.hu-berlin.de/de/content/architektur-und-morphogenese-von-biofilmen/
Gluconacetobacter aggeris Gluconacetobacter asukensis Gluconacetobacter azotocaptans Gluconacetobacter diazotrophicus Gluconacetobacter entanii Gluconacetobacter europaeus Gluconacetobacter hansenii Gluconacetobacter intermedius Gluconacetobacter johannae Gluconacetobacter kakiaceti Gluconacetobacter kombuchae Gluconacetobacter liquefaciens (type species of the genus) Gluconacetobacter maltaceti Gluconacetobacter medellinensis Gluconacetobacter nataicola Gluconacetobacter oboediens Gluconacetobacter rhaeticus Gluconacetobacter sacchari Gluconacetobacter saccharivorans Gluconacetobacter sucrofermentans Gluconacetobacter swingsii Gluconacetobacter takamatsuzukensis Gluconacetobacter tumulicola Gluconacetobacter tumulisoli Gluconacetobacter xylinus